Microscope Image Analysis
Automated 2D Segmentation and 3D Reconstruction of Neurites From Electron Microscope Images (Current Ph.D. Project)
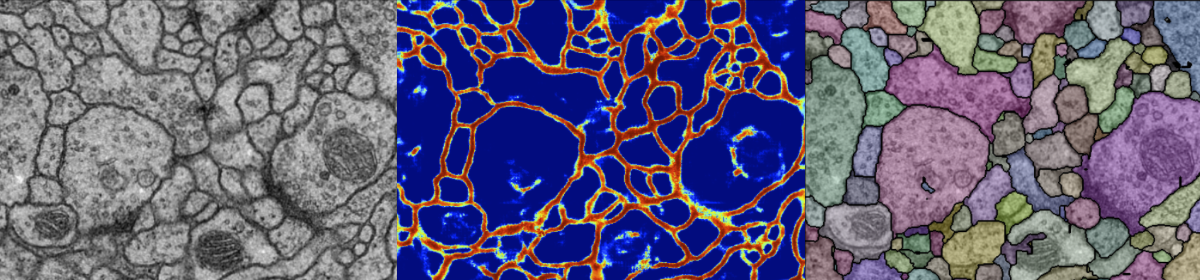
For automatic reconstruction of a micro-circuit of neurons from consecutive electron microscope images, firstly each cell membrane regions are detected by the classifier by using deep learning techniques (convolutional neural networks). Then applying the watershed algorithm, the initial over-segmentation hypothesis is obtained.
Virus Particle Detection by Convolutional Neural Network in Transmission Electron Microscopy Images
| Problem Settings | Prediction |
|---|---|

|

|
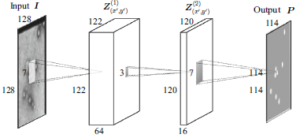
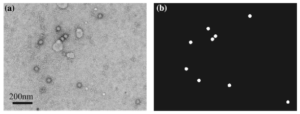
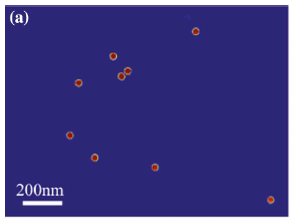
A new computational method for the detection of virus particles in transmission electron microscopy (TEM) images is presented. Our approach is to use a convolutional neural network that transforms a TEM image to a probabilistic map that indicates where virus particles exist in the image. Our proposed approach automatically and simultaneously learns both discriminative features and classifier for virus particle detection by machine learning, in contrast to existing methods that are based on handcrafted features that yield many false positives and require several postprocessing steps. The detection performance of the proposed method was assessed against a dataset of TEM images containing feline calicivirus particles and compared with several existing detection methods, and the state-of-the-art performance of the developed method for detecting virus was demonstrated. Since our method is based on supervised learning that requires both the input images and their corresponding annotations, it is basically used for detection of already-known viruses. However, the method is highly flexible, and the convolutional networks can adapt themselves to any virus particles by learning automatically from an annotated dataset.
- Eisuke Ito, Takaaki Sato, Daisuke Sano, Etsuko Utagawa, Tsuyoshi Kato, Virus Particle Detection by Convolutional Neural Network in Transmission Electron Microscopy Images. Food and environmental virology, Jan. 2018: 1-8. DOI: 10.1007/s12560-018-9335-7
Co-Localized Area Detection on Multi-Channel Fluorescent Microscope Images
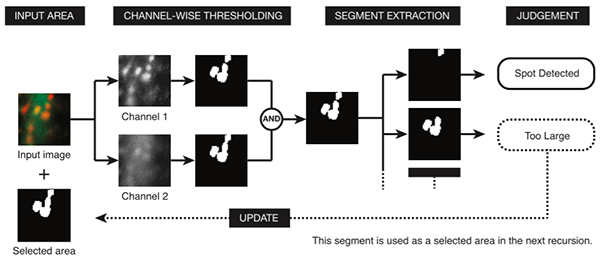
Automatic detection of immunoreactive areas in fluorescence microscopic images is becoming a key technique in the field of biology including neuroscience. In this study, we developed a new algorithm that exhaustively detects co-localized areas in multi-channel fluorescence images, where shapes of target objects may differ among channels. Different adaptive binarization thresholds for different local regions in different channels are introduced and the condition of each segment is assessed to recognize the target objects. The proposed method was applied to detect immunoreactive spots that labeled membrane receptors on dendritic spines of mouse cerebellar Purkinje cells.
- Eisuke Ito, Yusuke Tomaru, Akira Iizuka, Hirokazu Hirai and Tsuyoshi Kato, Adaptive Local Thresholding for Co-localization Detection in Multi-Channel Fluorescence Microscopic Images, IEICE Transactions on Information & Systems, Vol.E99-D,No.11,pp.-,Nov. 2016.
EEG Signal Classification
EEG Signal Classification on Manifolds
Multi-Class Motor-Imagery EEG Classification on Grassmann manifold and Riemannian manifolds. (Master thesis, 2014)